February 12, 2026 · Zifeng Wang
Paper: A foundation model for human-AI collaboration in medical literature mining (Nature Communications, 2025)

Starting from OpenScholar: the buzz around AI for literature synthesis
OpenScholar's recent appearance in Nature has given "using AI for scientific literature synthesis" another push: retrieval-augmented generation, 45 million open-access articles, citation accuracy on par with human experts, and generic LLMs' 78–90% citation hallucination rates are often cited for contrast [1]. Using large language models (LLMs) for literature mining and synthesis is already a trend that can't be ignored. Many of us have tried tools like ChatGPT for search and report generation. In my own experience, though, we're still far from something that's actually usable.
We'd like to ride that wave and introduce an earlier piece of work on medical literature mining and systematic review: LEADS, published in Nature Communications this year. Unlike OpenScholar's "open literature + general QA" angle [2], LEADS is built around the medical systematic-review pipeline: from search strategy and inclusion/exclusion screening to data extraction, all designed and evaluated around proprietary domain data and a fixed workflow in the medical setting.
Here we mainly want to make three points: the role of proprietary data, the importance of a domain-specific workflow in medicine, and the motivation and main results that follow.
Medical literature mining: why "generic model + open data" isn't enough
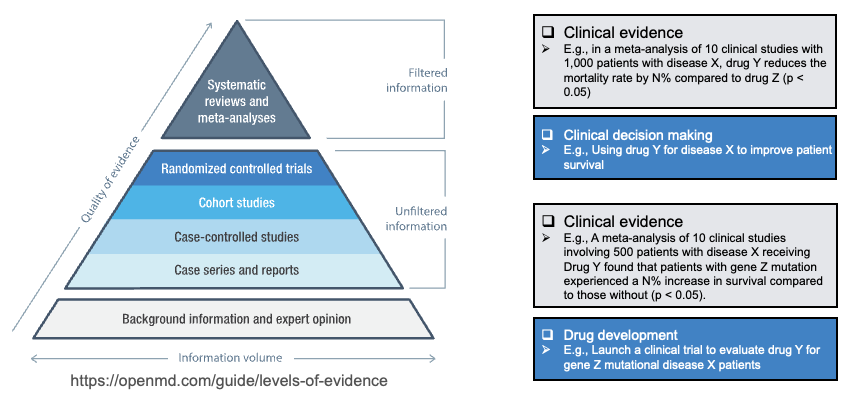
In medicine, literature mining, especially systematic review and meta-analysis, is a cornerstone of evidence-based practice: discovering, integrating, and interpreting new evidence, updating guidelines, and supporting industry and regulators. On the evidence pyramid, systematic reviews and meta-analyses sit at the top as "filtered" information; below them are single RCTs, cohort studies, and other "unfiltered" information. Doing literature mining and synthesis well directly affects the quality of clinical decisions and drug development (see figure below).
PubMed sees over 50,000 systematic reviews published per year [3,4], but a single review takes 67 weeks on average from start to finish [5], and top institutions and pharma spend tens of millions of dollars annually on literature review [6]. The literature is exploding, e.g. PubMed now indexes over 35 million records, with over a million new entries each year, and many entries have poor metadata, so search precision suffers [7], leading to incomplete searches, selection bias, and data-extraction errors in reviews [8].
Using a generic LLM (e.g. GPT-4o) for "search + read + synthesize" is one approach, but there are several hurdles:
- Task shape is different: medical systematic review follows a very fixed workflow: define PICO, generate search queries, search PubMed/ClinicalTrials.gov → title/abstract screening (citation screening) → full-text screening → data extraction (study characteristics, arms, participant counts, outcomes, etc.). These steps have clear inputs, outputs, and evaluation criteria; they're not covered by "ask a question and get a cited answer."
- Domain data is irreplaceable: what actually teaches a model "which search strategy retrieves the right studies," "what counts as eligible," and "how to pull PICO and results from full text" is large-scale existing systematic reviews and their cited literature, plus clinical trial registries. That data is proprietary and structured: which papers were included in which review, what each review's search strategy and extraction tables look like, and is barely present in open web or general corpora.
- Human–AI collaboration is the realistic path: literature synthesis demands high factual accuracy and traceability; full automation tends to hallucinate, miss studies, or extract wrongly. A more realistic route is: use AI to speed up search, screening, and extraction, with experts making final judgments and corrections. So we care not only about "model-vs-task metrics" but also "when experts work with LEADS, do they save time while keeping or improving recall/accuracy?"
So our motivation is straightforward: on this medical systematic-review workflow, train a foundation model for multiple subtasks using high-quality proprietary domain data, and validate in a human–AI collaboration setting whether it actually helps clinicians and researchers.
Proprietary data and LEADSInstruct: what we use and how
LEADS is trained and evaluated on linked data across systematic reviews, publications, and clinical trials:
- 21,335 systematic reviews from PubMed;
- 453,625 publication citations associated with those reviews;
- 8,485 of those reviews linked to 27,015 ClinicalTrials.gov trial records.
These data come from authoritative sources such as Cochrane (from PubMed), PubMed, and ClinicalTrials.gov; through a standardized workflow they are extracted and structured from raw articles and trial reports into tables and forms usable for training and evaluation (e.g. study characteristics, arm design, outcome data). The figure below summarizes this "data foundation": from proprietary sources on the left to structured papers, extraction tables, and trial data-entry forms on the right.
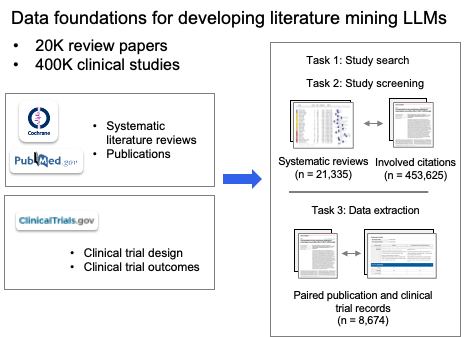
We split the three core pieces of systematic review methodology: literature search, citation screening, and data extraction, into six subtask types and built the instruction dataset LEADSInstruct (~633,759 instructions) from the above data, including: search-query generation (for publications and trials), study eligibility assessment, study characteristic extraction, participant statistics, arm design extraction, and trial result extraction. To our knowledge, the scale and task coverage make it the largest instruction-level benchmark for medical literature mining to date.
In one sentence: we don't use open web or general corpora for "generic reading"; we use "decisions and labels that review authors have already made" to train the model. Proprietary data here directly determines whether the model can learn the "rules" and details of medical systematic review.
Why the medical workflow deserves its own modeling
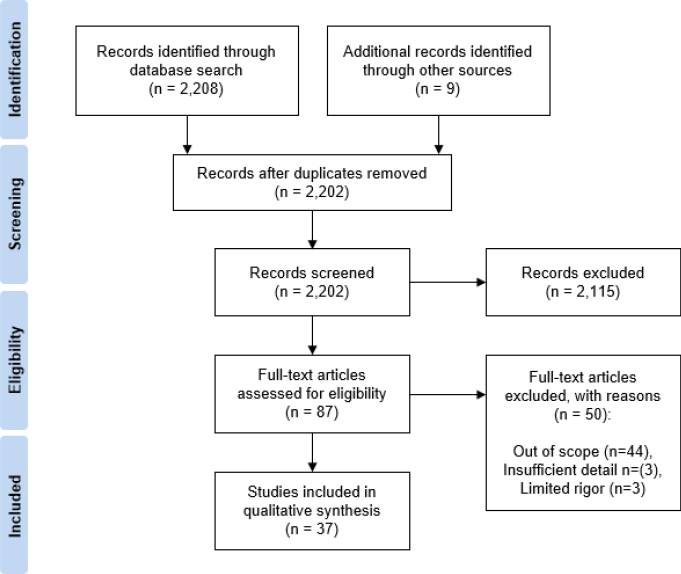
The systematic review process is highly standardized and usually reported via a PRISMA flow: Identification (search and deduplication) → Screening (title/abstract) → Eligibility (full-text eligibility) → Included (qualitative/quantitative synthesis). The figure below is a typical PRISMA example: from multi-database search yielding thousands of records, through dedup, title/abstract screening, and full-text assessment, down to dozens of studies included for qualitative or quantitative synthesis.
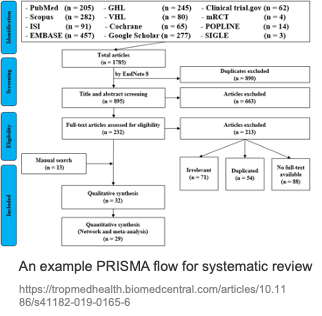
Each step has medical-specific details: search must produce queries that recall target studies while controlling noise on PubMed and ClinicalTrials.gov; screening must apply PICO and inclusion/exclusion criteria consistently across many titles/abstracts; data extraction must pull design, population, intervention, comparison, outcomes, and effect sizes from full text in a format suitable for meta-analysis. Traditionally this pipeline consumes a lot of labor and time; the human–AI idea is to introduce AI agents at key steps, with experts in the loop and AI accelerating, so that "many experts, many weeks" can be compressed toward "~1 expert, ~1 week" while preserving quality. The left side of the next figure sketches this AI-augmented workflow (experts working with multiple AI agents to produce research questions and clinical evidence); the right side shows the full PRISMA flow for comparison.
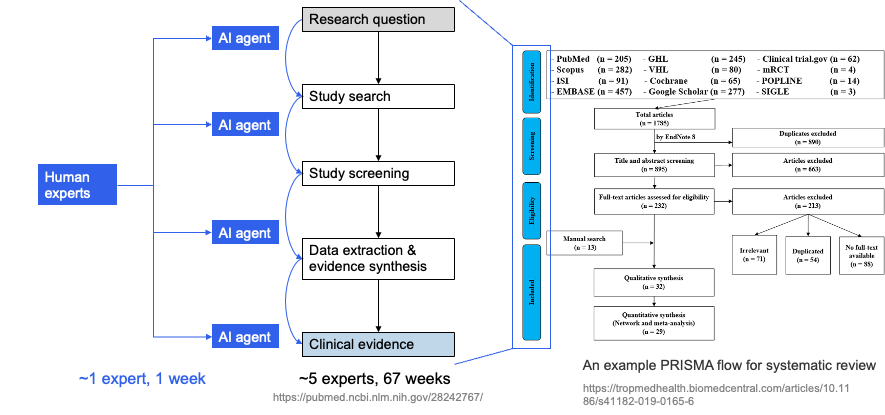
These steps are chained; doing "one step" well isn't enough—the model needs to handle multiple steps in a single framework and adapt to different review topics. LEADS explicitly decomposes this full workflow into six subtask types and uses one LEADSInstruct-based instruction tuning so a single model can do multi-task, multi-topic work without retraining for each new review.
How LEADS performs: ahead of generic LLMs and real time savings
We ran offline evaluation on thousands of systematic reviews and hundreds of thousands of studies, comparing against GPT-4o, GPT-3.5, Haiku, Mistral, Llama, and medical-domain models such as BioMistral and MedAlpaca; we also ran a pseudo-prospective evaluation (31 reviews published after 2025) to reduce data-leakage concerns.
At the offline task level (see paper Fig. 1d, Fig. 2–4): LEADS outperforms all compared models across the six task types. For search-query generation, LEADS' recall is a clear step above the best baseline (publication search +3.76, trial search +7.43); for screening and the various extraction tasks, LEADS leads on Recall@50 or Accuracy as well. Using generic Mistral-7B as the base and only instruction tuning on LEADSInstruct already yields a large gain over zero-shot GPT-4o, showing that domain instruction data alone can substantially raise performance on medical literature mining. The figure below summarizes LEADS vs. GPT-3.5, Haiku-3, GPT-4o, and Mistral-7B on nine subtasks (search-query generation, study eligibility, study characteristics / arm design / participant stats / trial result extraction, etc.): LEADS' radar plot extends clearly outward on most dimensions, especially for study eligibility and the extraction tasks.
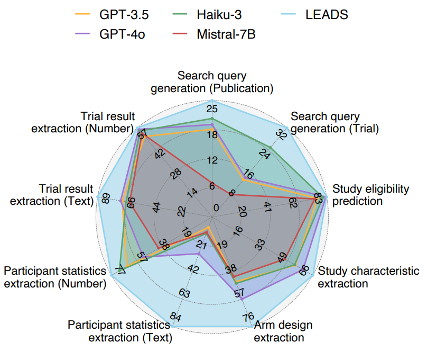
On the human–AI side: we ran a user study with 16 clinicians and researchers from 14 institutions, comparing "expert only" vs. "expert + LEADS":
- Citation screening: with LEADS, recall 0.81 vs. 0.78 (without LEADS), with ~20.8% time saved;
- Data extraction: accuracy 0.85 vs. 0.80, ~26.9% time saved.
So LEADS doesn't replace experts; it shortens screening and extraction time while maintaining or improving quality. That's a direct answer to the "reviews are too slow and expensive" pain point.
Space limits us to a subset of results here; for more task breakdowns, ablations, and pseudo-prospective experiments, see the paper.
Summary: proprietary data + domain workflow + human–AI collaboration
OpenScholar showed the potential of open scientific literature + retrieval augmentation + controlled citations for general scientific QA; LEADS shows what proprietary data and workflow modeling for search, screening, and extraction can do on the more vertical, process- and evidence-conscious track of medical systematic review:
- Proprietary data: LEADSInstruct built from 20k+ systematic reviews, 450k+ publications, and 27k+ trial registry records lets the model learn "how medical reviews are done" instead of staying at generic reading.
- Domain workflow: breaking search, screening, and extraction into trainable, evaluable subtasks and handling them in one foundation model is what allows both broad topic coverage and alignment with real workflows.
- Human–AI collaboration: in our user study, experts using LEADS saved time and improved recall/accuracy on screening and extraction, showing that "AI accelerates, experts gatekeep" is a practical and deployable direction today.
We believe that in literature mining and synthesis, vertical proprietary data and workflow-specific modeling will matter more and more; LEADS is our systematic step on the medical track, and we welcome discussion and collaboration with anyone working on systematic review, evidence-based medicine, or medical NLP.
References
[1] OpenScholar: Synthesizing scientific literature with retrieval-augmented language models. Nature (2026). nature.com/articles/s41586-025-10072-4
[2] Wang, Z. et al. A foundation model for human-AI collaboration in medical literature mining. Nat. Commun. 16, 8361 (2025). doi.org/10.1038/s41467-025-62058-5
[3] PubMed systematic review statistics (e.g. annual growth).
[4] Bastian, H., Glasziou, P. & Chalmers, I. Seventy-five trials and eleven systematic reviews a day: how will we ever keep up? PLoS Med. 7, e1000326 (2010).
[5] Borah, R. et al. Analysis of the time and workers needed to conduct systematic reviews of medical interventions using data from the PROSPERO registry. BMJ Open 7, e012545 (2017).
[6] See LEADS paper for citations on NIH-funded institutions' and pharma's literature review costs.
[7] See LEADS paper for discussion of PubMed metadata and search precision.
[8] See LEADS paper for review of common issues in systematic reviews (e.g. incomplete search, selection bias, extraction errors).