February 14, 2026 · Zifeng Wang
Paper: Nature Biomedical Engineering · Code & data: GitHub · Platform demo: biodsa.github.io

Building AI Agents for Automated Research
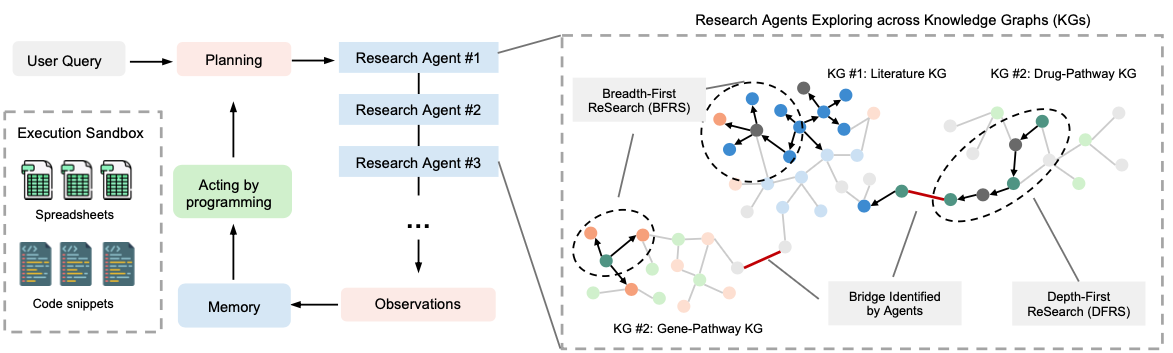
AI for Science has been in the spotlight in recent years. The potential of large language models (LLMs) and AI agents is being explored in many directions. Earlier work includes the 2023 Nature paper on Coscientist [1], which showed how LLMs can be used for document search, code execution, and experiment automation; Google's AI co-scientist system [2], released in early 2025, uses multiple specialized agents to generate, validate, and refine scientific hypotheses, with applications in drug repurposing, target discovery, and antimicrobial resistance.
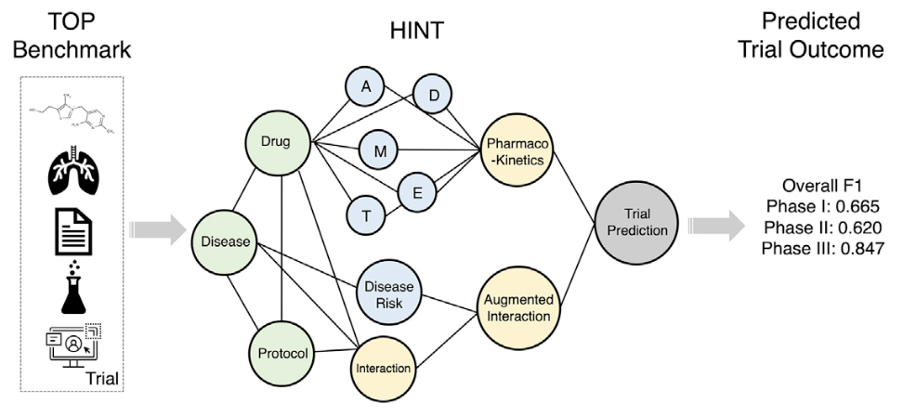
I have been using LLMs to support my own research for a while. In 2023, I mainly relied on GPT to write small scripts, but often after many rounds of prompting the code was still unusable. Starting in 2024, at Keiji AI, we began to systematically study code-generating AI agents for drug discovery, which largely involves bioinformatics workflows. Those workflows can be more complex than writing generic scripts: integrating multiple omics data, running non-trivial statistical tests, and producing standard visualizations such as Kaplan–Meier survival curves, OncoPrint heatmaps, and pathway enrichment analyses. In these steps, LLMs are especially prone to mistakes in data handling, and a single error can invalidate the whole analysis.
So from the outset we focused on how to systematically evaluate existing LLMs' ability to perform bioinformatics-style analysis. We first looked at existing benchmarks, such as HumanEval [3], DS-1000 [4], and SWE-Bench [5], which target general programming or software engineering and do not cover the specific needs of biomedical data science. BioCoder [6] covers code generation for some bioinformatics tools but does not include many end-to-end analysis tasks.
BioDSBench: Building a Coding Benchmark from Published Biomedical Papers
We reasoned that the figures and tables in a paper largely reflect the methods and results of the analyses. If we turn those figures and tables into programming tasks and ask LLMs to complete them, can we assess how well LLMs perform at bioinformatics-style analysis?
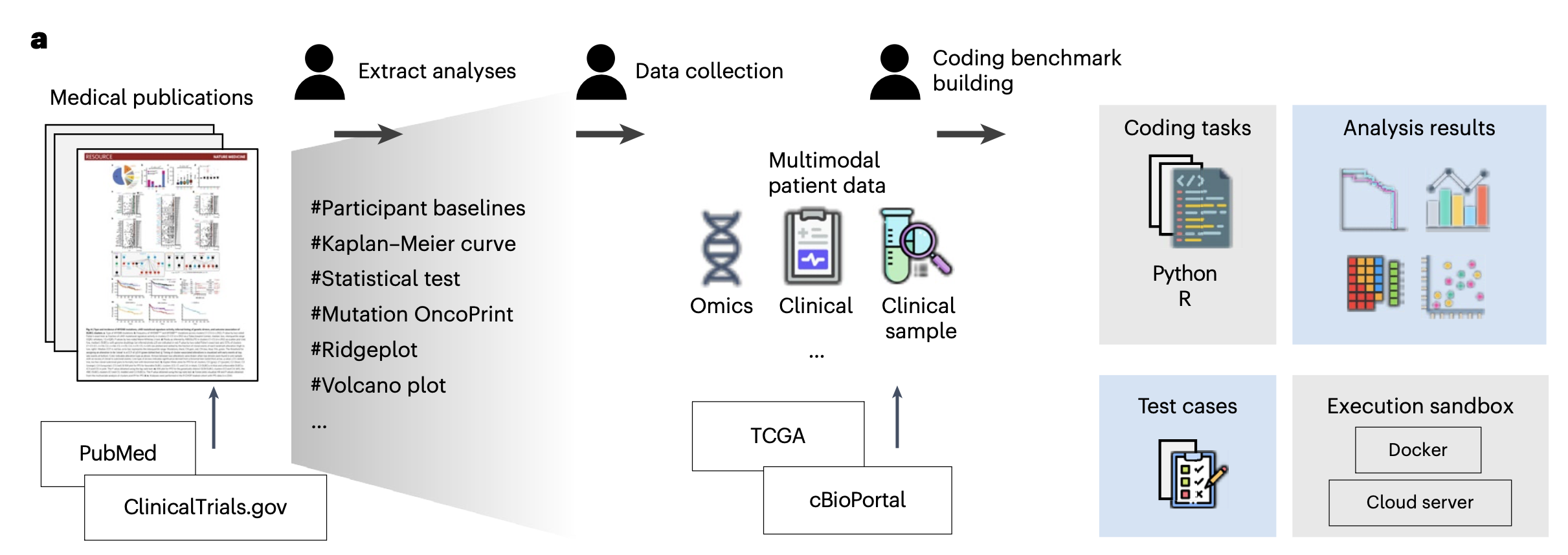
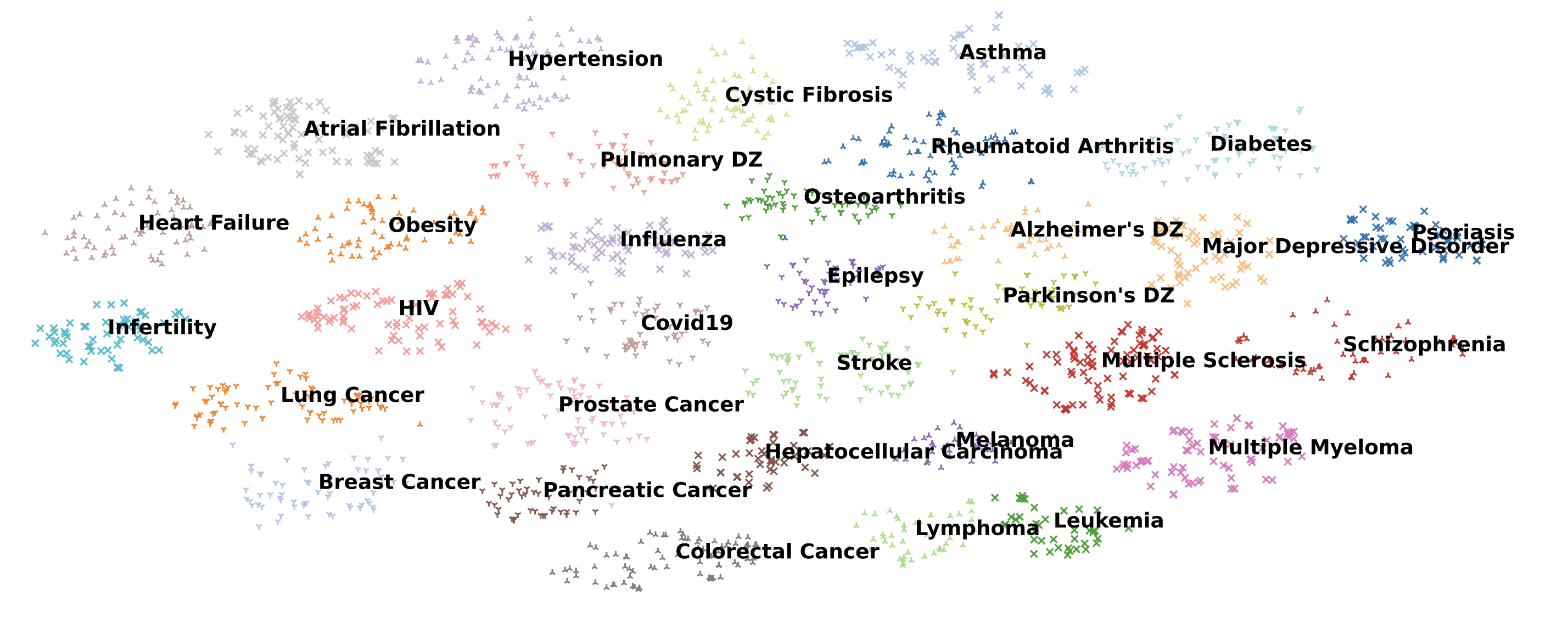
Concretely, we selected 39 biomedical studies with available raw patient data from cBioPortal, spanning seven research areas: biomarker discovery, integrative analysis, genomic profiling, molecular characterization, therapeutic response, translational research, and pan-cancer analysis. From these papers we extracted the analyses actually performed by the authors and grouped them into eight analysis types: gene expression, genomic alteration, survival analysis, clinical feature engineering, treatment response, descriptive statistics, enrichment and pathway analysis, and data integration. We then built a benchmark of 293 programming tasks in both Python and R.
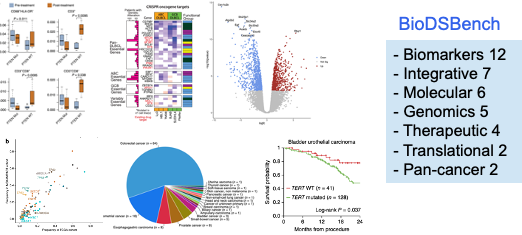
Each task has a clear structure: an input question, a dataset schema description, prefix code (simulating the context of a notebook), reference code, and automated test cases. Importantly, our benchmark does not only check whether the code runs; it checks whether the outputs of the LLM-generated code match the results reported in the original papers.
That distinction matters. In general programming benchmarks, tests usually check function inputs and outputs. In data science, code that runs without error can still be wrong. A survival analysis script might execute without any exception but use the wrong grouping variable, so that the resulting Kaplan–Meier curves differ completely from those in the paper.
How LLMs Perform on Biomedical Data Science: Not Yet There
We evaluated 8 proprietary models (GPT-4o, GPT-4o-mini, Gemini-Pro, Gemini-Flash, Opus-3, Sonnet-3.5, o3-mini, DeepSeek-R1) and 8 open-source models (e.g. Llama-3-8B/70B, CodeLlama-7B/34B, StarCoder2-15B, Qwen-Coder-7B/32B) under multiple prompting strategies.
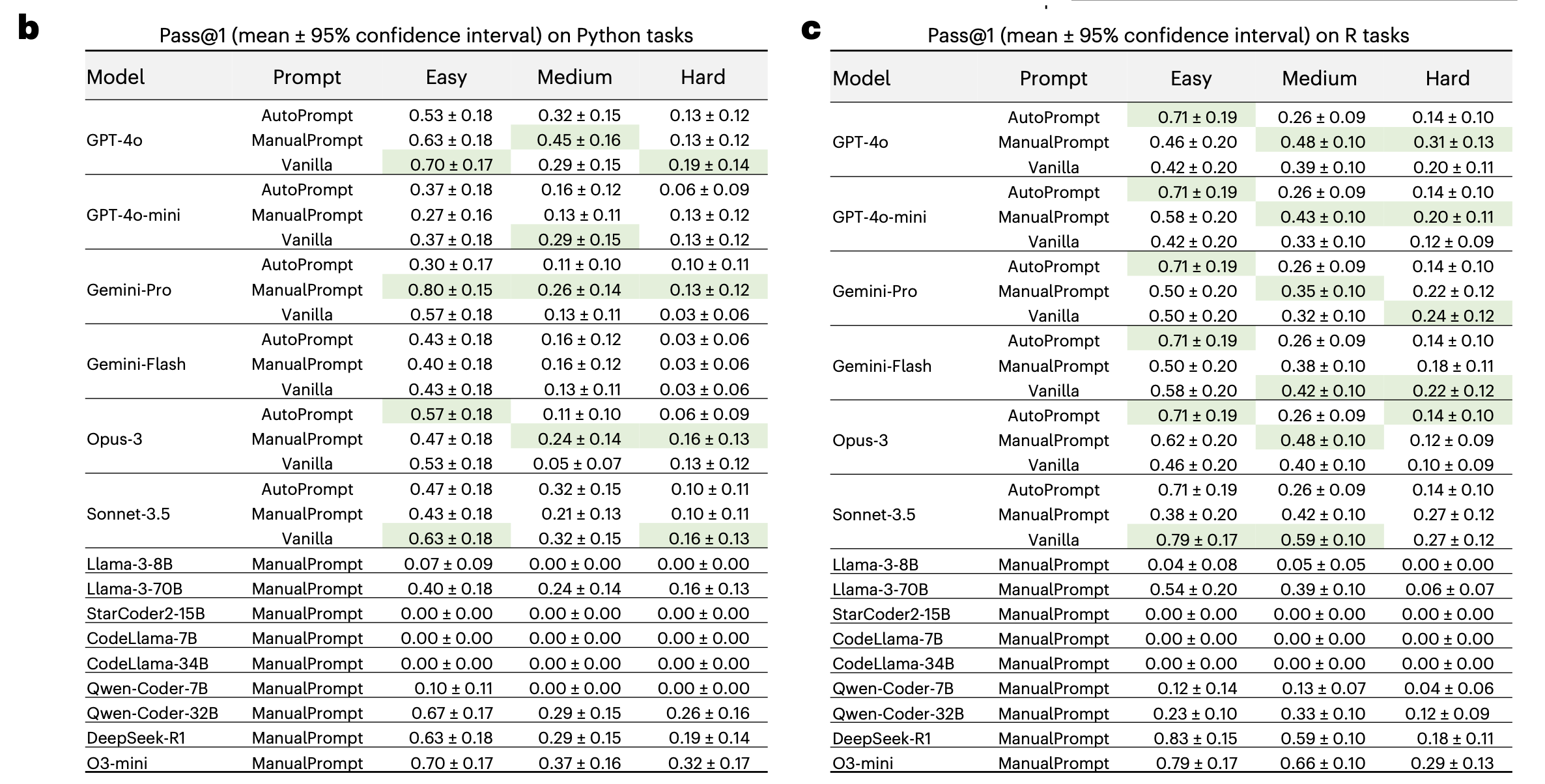
The results were in line with what we expected. On Python tasks, accuracy ranged from 40–80% on easy, 15–40% on medium, and 5–32% on hard; R showed a similar pattern. Even the best models, o3-mini and DeepSeek-R1, reached only about 30% on hard tasks.
When we looked at failure modes, we found that the most common was not runtime crashes but "test failure": the code runs to completion but produces incorrect analysis results. That is especially risky in biomedical settings, where domain experts with little programming experience may assume that code that runs is correct.
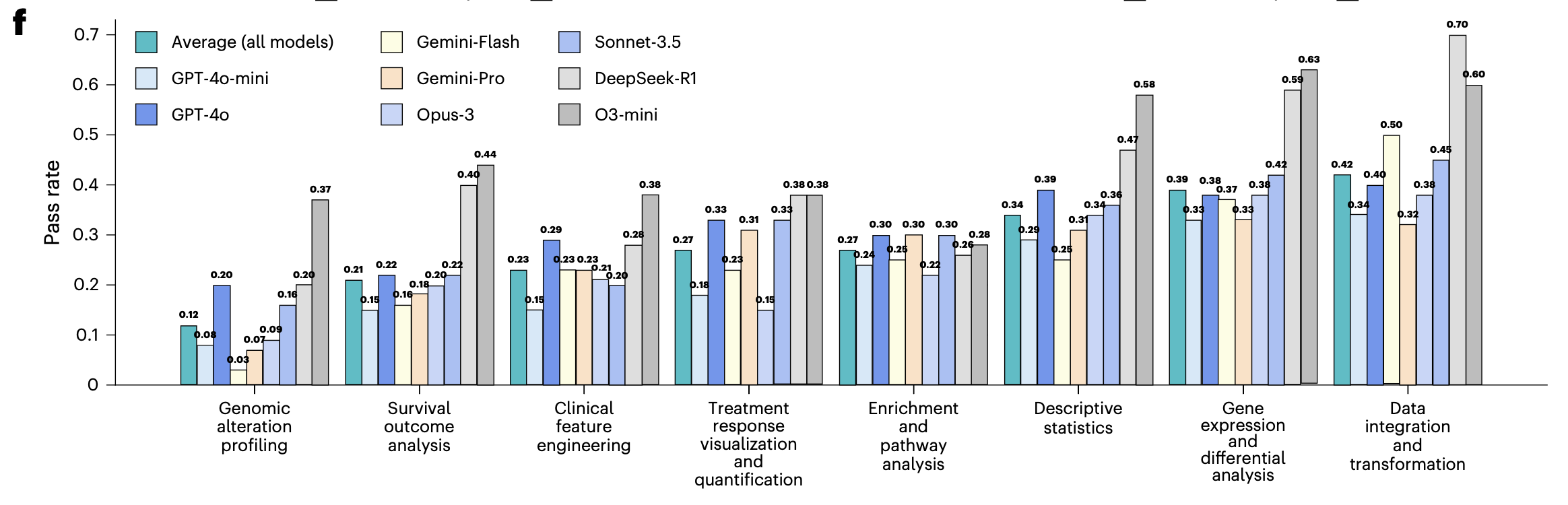
By analysis type, genomic alteration profiling (mean 0.12) and survival outcome analysis (mean 0.21) were the hardest; data integration and transformation (mean 0.42) was relatively easier. Reasoning-oriented models such as o3-mini and DeepSeek-R1 did better than average across task types, but absolute performance still has room to improve.
Main Finding: The Analysis Plan Is Key to Success
We manually inspected many failure cases and identified a core issue: when given a data analysis request, LLMs show high variability in how they interpret the task and the structure of the data. The same question can lead to very different analysis paths across models, or even across runs of the same model. To improve correctness, we asked whether providing a structured analysis plan before code generation—spelling out what to do and how—would significantly improve LLM performance. That is a natural idea, since human scientists also plan their analysis steps before coding.
The answer is yes. When we gave the LLM a human-written analysis plan as extra input (we call this PlanPrompt), accuracy on medium and hard tasks improved by 0.13 and 0.35 over the best other prompting methods. So the analysis plan is the cornerstone of reliable data science programming.
DSWizard: An Agent Framework Centered on Analysis Plan and Reasoning
Based on that idea we designed the DSWizard agent. Unlike a traditional CoderAgent (ReAct-style, with the LLM freely exploring data and iterating on code), DSWizard is built around "plan first, then code."
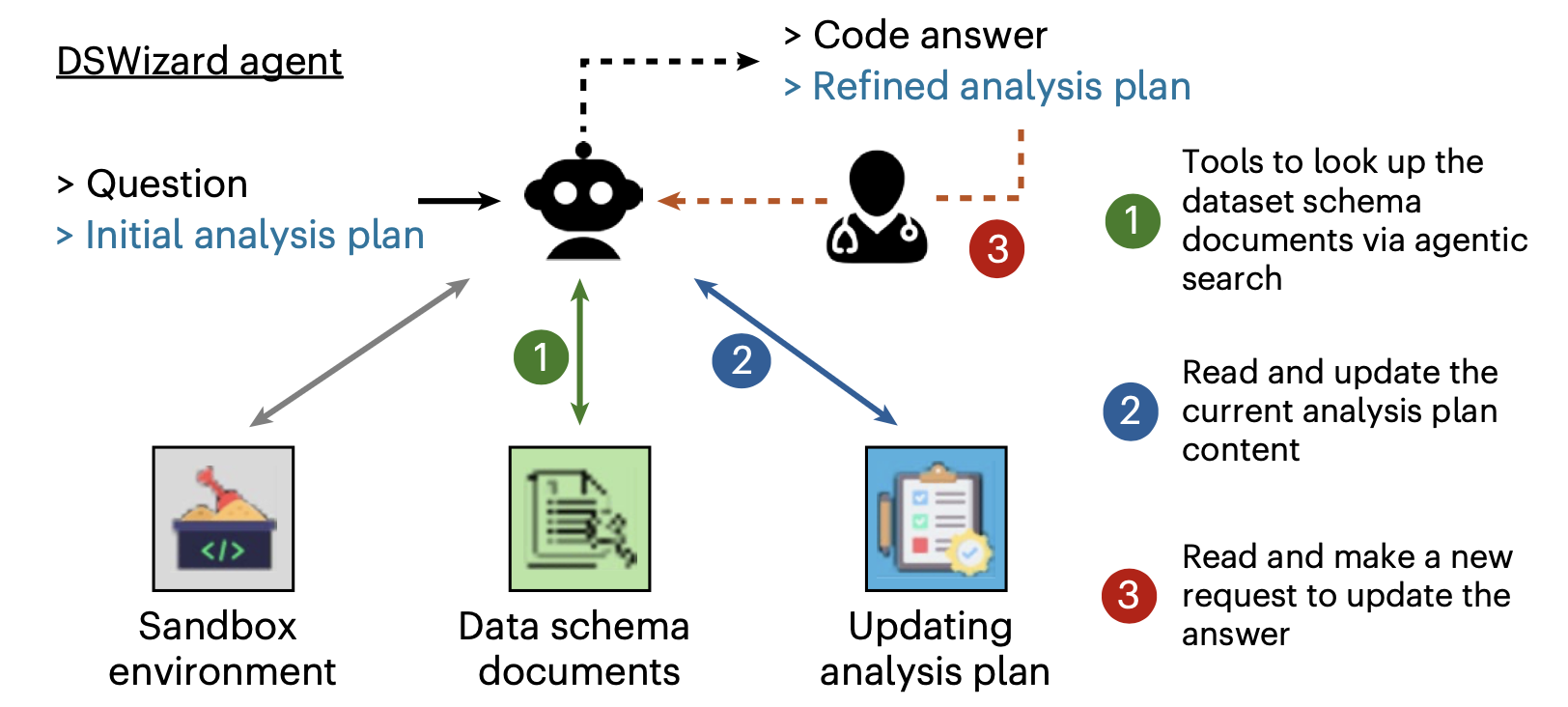
DSWizard has three main components: (1) a data schema index so the agent can query table structure, column names, and types without writing exploratory code; (2) a structured analysis plan as input that defines each step; and (3) a plan-update loop where the agent refines the plan while exploring the data and only then turns the final plan into code.
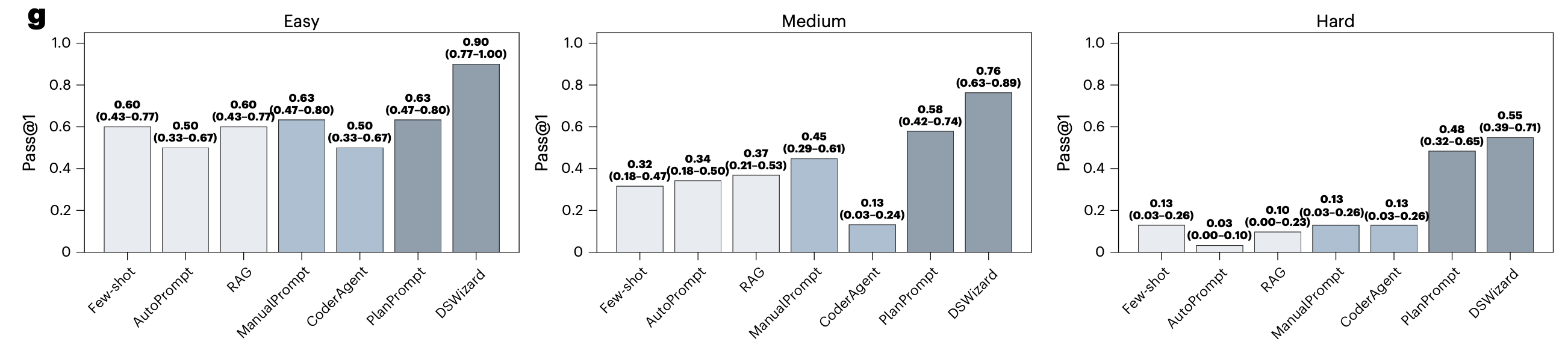
The results are clear: DSWizard reached 0.74 average accuracy and outperformed Vanilla prompting on all 11 held-out studies. By difficulty, it reached 90% on easy, 76% on medium, and 55% on hard, with gains of 27%, 31%, and 42% over the best non-planning baseline. Interestingly, giving the LLM a flexible CoderAgent (ReAct-style) did worse than single-round Vanilla prompting. Unconstrained exploration hurts: the agent either wastes steps on unproductive data exploration or commits too early to a wrong answer. A more constrained, plan-driven agent does better.
In retrospect, our work on planning was an attempt to use test-time reasoning to address the limitations of LLMs on data science tasks. In September 2024, OpenAI released the o1 family, marking the rise of reasoning models—consistent with what we had observed earlier that year. We were trying to elicit stronger reasoning from non-reasoning models. In rebuttal we added several reasoning models, including o3-mini and DeepSeek-R1, and they did show substantial gains on our benchmark.
From Benchmark to Platform: Human–AI Collaboration in Data Science
With the benchmark and agent framework in place we built a usable platform. The idea is that users and the AI jointly build an analysis plan, check that the tables and columns referenced in the plan exist in the data, and then turn the plan into executable code. The platform includes a sandbox so users can run code and inspect results directly.
We recruited five medical researchers for a user study. Each tried to reproduce analyses from three bioinformatics papers, with ten tasks per paper. On easy tasks, 84–88% of the code in their final submissions came directly from LLM-generated code; the share was lower on medium and hard tasks, but overall the platform substantially improved the efficiency of domain experts with limited programming experience. In follow-up discussions we summarized four practical guidelines:
- Plan before code: build a concrete analysis plan with the AI instead of asking it to "analyze the data for me."
- Validate the data schema: confirm that the tables and columns referenced in the plan actually exist.
- Provide context: explain domain-specific concepts (e.g. how "overall response" is defined).
- Code that runs is not necessarily correct: review and revise the analysis plan after inspecting the results.
Application: AI Agents for Automated Biomarker Discovery
A natural extension of the DSWizard framework is to let an AI agent carry out a full biomedical analysis pipeline—for example, biomarker discovery from data.
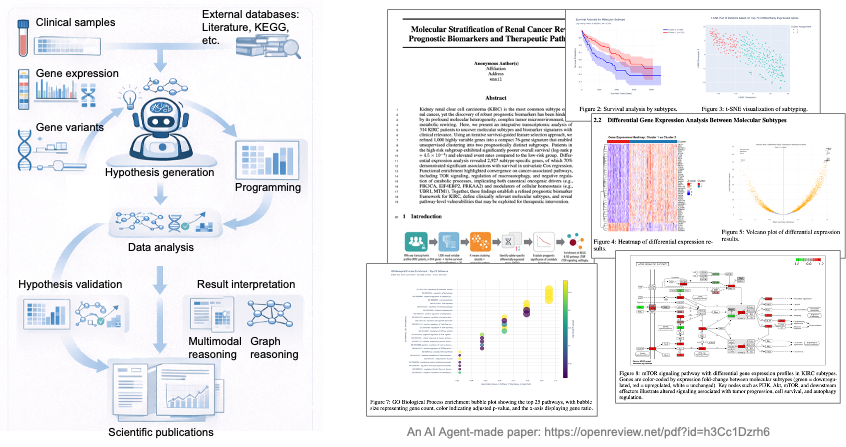
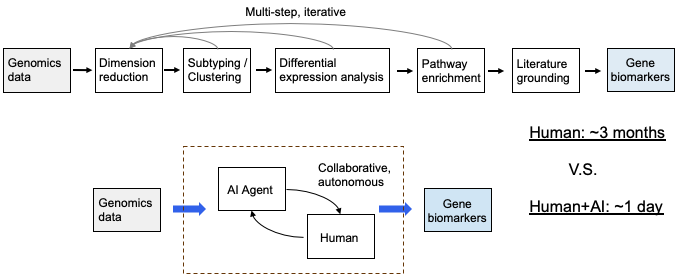
A recent piece of work [7] explores this direction. We use an AI agent to take in genomics data and automatically run a pipeline: dimensionality reduction, clustering, differential expression, survival analysis, pathway enrichment, and finally biomarker discovery with prognostic relevance. In one example, our agent worked on the kidney renal clear cell carcinoma (KIRC) dataset. Starting from transcriptomic data for 314 patients, it used iterative survival-guided feature selection to obtain a 76-gene signature that split patients into two molecular subtypes with significantly different prognosis (log-rank p = 4.5×10⁻⁴). Differential expression analysis then identified 2,927 subtype-specific genes, 70% of which were significantly associated with overall survival in univariate Cox regression. Pathway enrichment highlighted cancer-related pathways such as TOR signaling, macroautophagy regulation, and negative regulation of catabolic processes. A paper generated by our agent is available here: openreview.net/pdf?id=h3Cc1Dzrh6
After talking with our bioinformatics colleagues we learned that the same pipeline could take months when done manually. With an AI agent, going from data to results can be done in a day. That does not mean the AI can replace human judgment—hypotheses and results from the agent still need to be validated and interpreted by domain experts. But it shows the potential of AI as a data co-scientist: greatly speeding up the analysis loop so that researchers can focus more on the science and less on implementation.
Summary
In short: LLMs are not yet fully reliable for biomedical data analysis, but an agent framework built around analysis plans can substantially narrow the gap.
BioDSBench provides a testbed for systematically evaluating LLM programming ability; DSWizard demonstrates the value of a plan-driven approach for human–AI collaboration in data science; and BioDSA explores how far AI agents can go on harder biomarker discovery tasks. From benchmark to agent to application, we hope this thread offers useful steps toward reliable AI data co-scientists.
Code and data are public: the benchmark is on HuggingFace and the code on GitHub. We welcome feedback and discussion.
References
[1] Boiko, D. A. et al. Autonomous chemical research with large language models. Nature 624, 570–578 (2023). nature.com/articles/s41586-023-06792-0
[2] Google Research. Accelerating scientific breakthroughs with an AI co-scientist. research.google/blog
[3] Chen, M. et al. Evaluating Large Language Models Trained on Code. arxiv.org/abs/2107.03374
[4] Lai, Y. et al. DS-1000: A Natural and Reliable Benchmark for Data Science Code Generation. In ICML 2023. arxiv.org/abs/2211.11501
[5] Jimenez, C. E. et al. SWE-bench: Can Language Models Resolve Real-World GitHub Issues? In ICLR 2024. arxiv.org/abs/2310.06770
[6] Tang, X. et al. BioCoder: A Benchmark for Bioinformatics Code Generation with Large Language Models. Bioinformatics, 2024. arxiv.org/abs/2308.16458
[7] Wang, Z., Danek, B. & Sun, J. BioDSA-1K: Benchmarking Data Science Agents for Biomedical Research. arxiv.org/abs/2505.16100
[8] Wang, Z. et al. Making large language models reliable data science programming copilots for biomedical research. Nature Biomedical Engineering, 2026. doi.org/10.1038/s41551-025-01587-2